DeeplyGrad Part 2: Teaching Our Autograd to Classify MNIST Digits
In Part 1 of the series, we built a Tensor class with full backpropagation support and trained a linear regression model. In this post, we add everything needed to train a real neural network. We will build PyTorch like Module for layers, activations, losses. Then we will put everything together to train a MLP classifier to classify the classic MNIST digits with high accuracy.
Code for this series is available on GitHub
Everything is a Module
In PyTorch, nn.Linear, nn.ReLU, nn.CrossEntropyLoss are Modules. There’s a reason PyTorch chose this design. When every component, a linear layer, an activation, a loss function, even an entire GPT model, is the same type, they snap together like LEGO blocks. You can plug any Module into any other Module, nest them arbitrarily deep, and the framework handles the rest. That uniformity is what makes it possible to build complex architectures from simple, reusable pieces.
Let’s build this abstraction first, then use it for everything else.
"""
deeplygrad.nn: Neural network building blocks
"""
from deeplygrad.tensor import Tensor
from typing import Optional
import numpy as np
from deeplygrad.backend import xp
class Module:
"""
Base class for all neural network components
This gives each component a uniform interface:
- forward() defines the computation
- parameters() collects all the trainable parameters
- zero_grad() resets the gradients of all the parameters of the module
- train()/eval() toggles the training mode
- register_buffer() stores non-learnable states
Subclass inherits and implements the __init__and forward.
"""
def __init__(self):
self.training: bool = False
self._buffers: Dict[str, Optional[Tensor]] = {}
def parameters(self):
"""Recursively collect all Tensors that require grad."""
buffer_names = set(self._buffers.keys()) if hasattr(self, '_buffers') else set()
params = []
for attr_name in dir(self):
if attr_name.startswith('_'):
continue
if attr_name in buffer_names:
continue
attr = getattr(self, attr_name)
if isinstance(attr, Tensor) and attr.requires_grad:
params.append(attr)
elif isinstance(attr, Module):
params.extend(attr.parameters())
elif isinstance(attr, (list, tuple)):
for item in attr:
if isinstance(item, Module):
params.extend(item.parameters())
elif isinstance(attr, dict):
for item in attr.values():
if isinstance(item, Module):
params.extend(item.parameters())
return params
def zero_grad(self):
"""Zero all parameter gradients in this module and it's children"""
for param in self.parameters():
param.zero_grad()
def register_buffer(self, name: str, tensor: Optional[Tensor]):
"""
Registers a non-learnable tensor as part of the module's state
Buffers are tensors which:
- Are not parameters (won't be returned by parameters())
- Are part of module's state (e.g. causal masks, running stats)
- Are accesible as self.name
Example (we'll do it when we'll implement transformer causal self-attention)
mask = Tensor(np.tril(np.ones(n_ctx, n_ctx)))
self.register_buffer('mask', mask)
"""
self._buffers[name] = tensor
setattr(self, name, tensor)
def train(self, mode: bool=True) -> 'Module':
"""
Sets the training mode. Affects behavior of Dropout, BatchNorm etc
Recurses into child modules
Usage:
model.train()
model.eval()
"""
self.training = mode
for attr_name in dir(self):
if attr_name.startswith('_'):
continue
attr = getattr(self, attr_name)
if isinstance(attr, Module):
attr.train(mode)
return self
def eval(self) -> 'Module':
return self.train(mode=False)
def __call__(self, *args, **kwargs):
return self.forward(*args, **kwargs)
def forward(self, *args, **kwargs):
raise NotImplementedError
The parameters() method walks through all attributes, finds any Tensor with requires_grad=True, and recurses into child Modules. It skips registered buffers (non-learnable state like causal masks, which we’ll use in Part 3). When we later build an MLP that contains Linear layers and ReLU activations, a single call to model.parameters() collects every trainable weight and bias automatically.
We also set up training mode and register_buffer() now. The training flag matters for layers like Dropout and BatchNorm that behave differently during training and inference. register_buffer() stores tensors that are part of the module’s state but should not be treated as learnable parameters. We’ll use both of these in Part 3 when we build the transformer.
Note that there’s no backward() on Module. That’s intentional. The computation graph lives in the Tensors, not in the Modules. When forward() runs, it creates Tensors through operations (@, +, relu, etc.), and each operation registers its _grad_fn on the output Tensor. By the time forward() returns, the graph already exists. So loss.backward() walks the Tensor graph directly. The Module is just an organizer for the forward pass.
Now everything we build inherits from Module. Subclasses call super().__init__() and implement forward(). That’s it.
Linear Layer
The most fundamental module, a fully connected layer: $\text{out} = x @ W+b$
"""
deeplygrad.nn: Neural network building blocks
"""
class Linear(Module):
def __init__(self, in_features: int, out_features: int, bias: bool=True):
super().__init__()
scale = np.sqrt(2.0 / in_features)
self.weight = Tensor(
np.random.randn(in_features, out_features) * scale,
requires_grad=True)
self.bias = Tensor(np.zeros(out_features), requires_grad=True) if bias else None
def forward(self, x: Tensor) -> Tensor:
out = x @ self.weight
if self.bias is not None:
out = out + self.bias
return out
We use Kaiming (He) initialization, scaling weights by $\sqrt{2/\text{fan_in}}$. This keeps the variance of activations roughly constant through ReLU layers. Bad initialization can make training painfully slow or prevent convergence entirely.
Notice that forward() uses the @ and + operators from our Tensor class. The autograd graph builds itself automatically as these ops execute.
Why Do We Need Nonlinearity?
Before implementing activation functions, let’s understand why they matter. A linear layer computes $y=xW + b$. If we stack two linear layers:
\[y = (xW_1 + b_1)W_2 + b_2 = xW_1W_2 + b_1W_2 + b_2\]This is just another linear function. No matter how many linear layers we stack, the result is always linear. The network can’t learn curves, decision boundaries, or anything more complex than a hyperplane.
Activation functions break this linearity. By inserting a nonlinear function between layers, each layer can bend the space in a different way. This is what gives deep networks their expressive power.
Activation Functions: ReLU and GELU
Since everything is a Module, our activations are Modules too.
ReLU
\(\text{ReLU}(x) = \text{max}(0, x)\)
The simplest nonlinearity. It clips negative values to zero. The gradient is 1 where input is positive, 0 where it’s negative. In Part 1, every Tensor operation (add, mul, matmul, etc.) needed a hand-written _backward because we wanted results to propagate for these elementary operation in the backward pass. But now that we have those primitives, Modules can compose them and get backward for free. The autograd engine handles the chain rule automatically.
ReLU is a perfect example. We added comparison operators (>, <, etc.) and a differentiable where() to our Tensor in Part 1. So ReLU is just:
"""
deeplygrad.nn: Neural network building blocks
"""
class ReLU(Module):
def __init__(self):
super().__init__()
def forward(self, t: Tensor) -> Tensor:
return where(t > 0, t, Tensor(0.0))
Notice no custom _backward. The autograd composes the backward passes of > and where() automatically via the chain rule. This is exactly how PyTorch modules work: if your forward() only uses existing differentiable ops, backward is free.
GELU
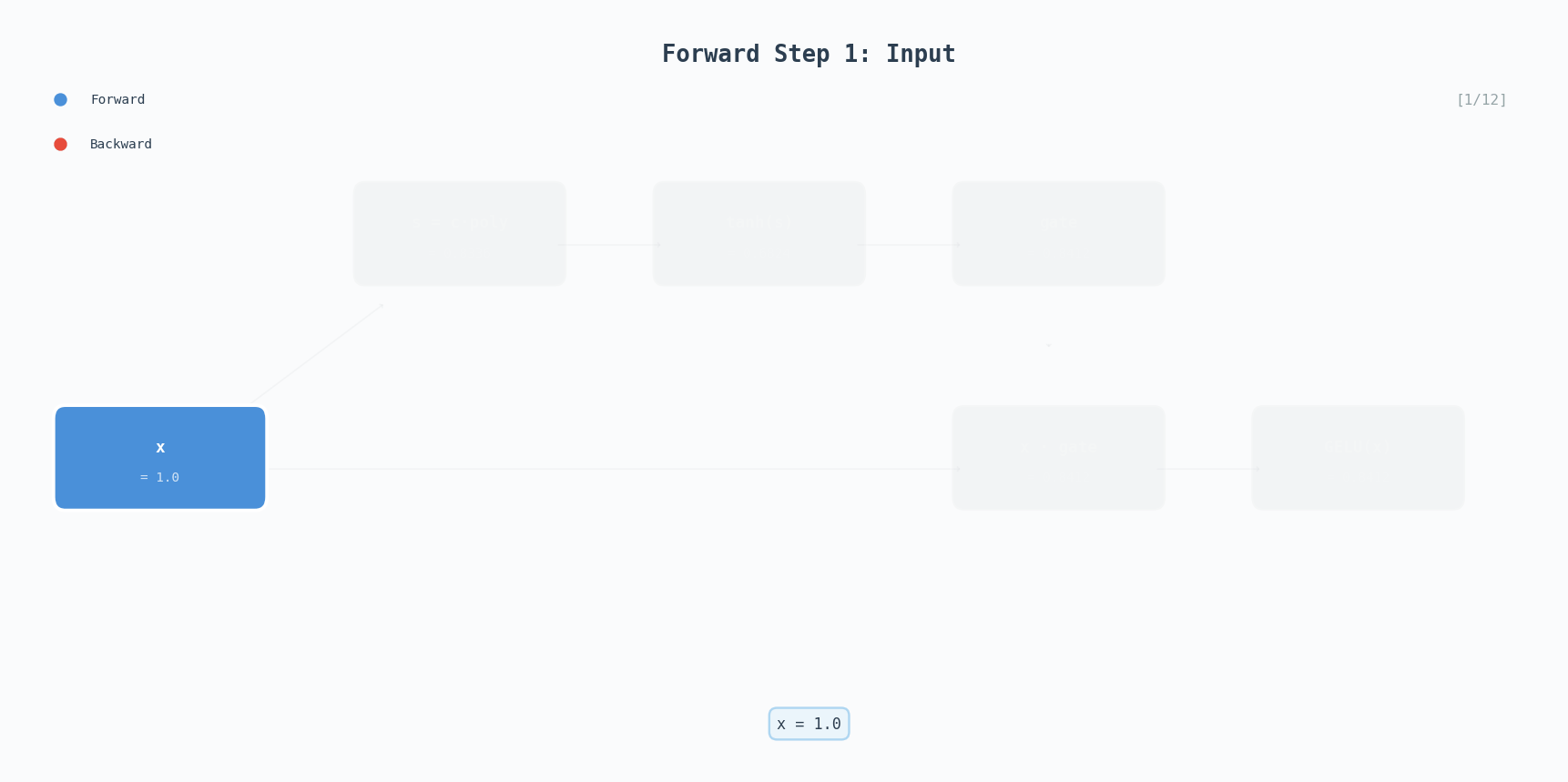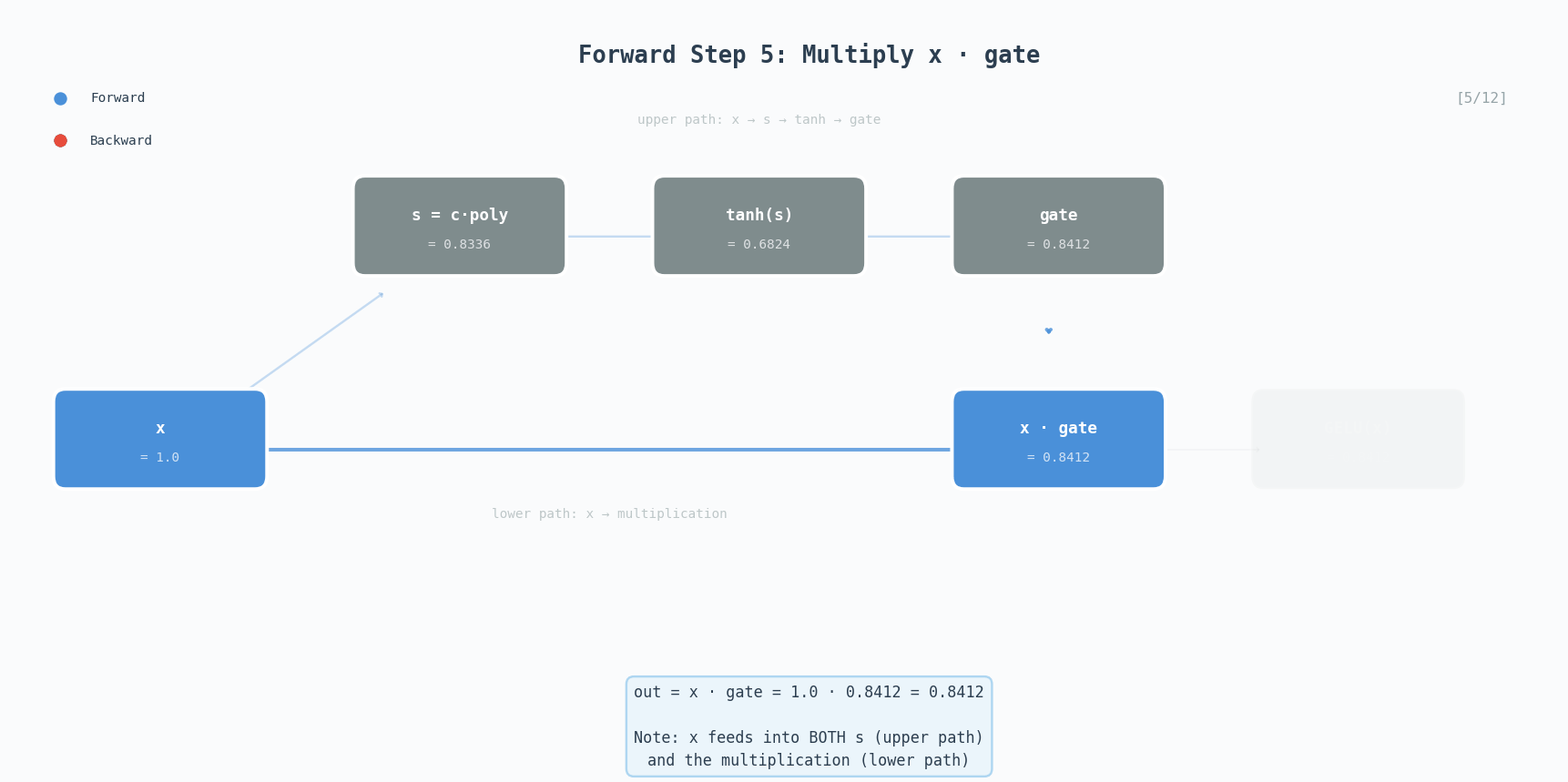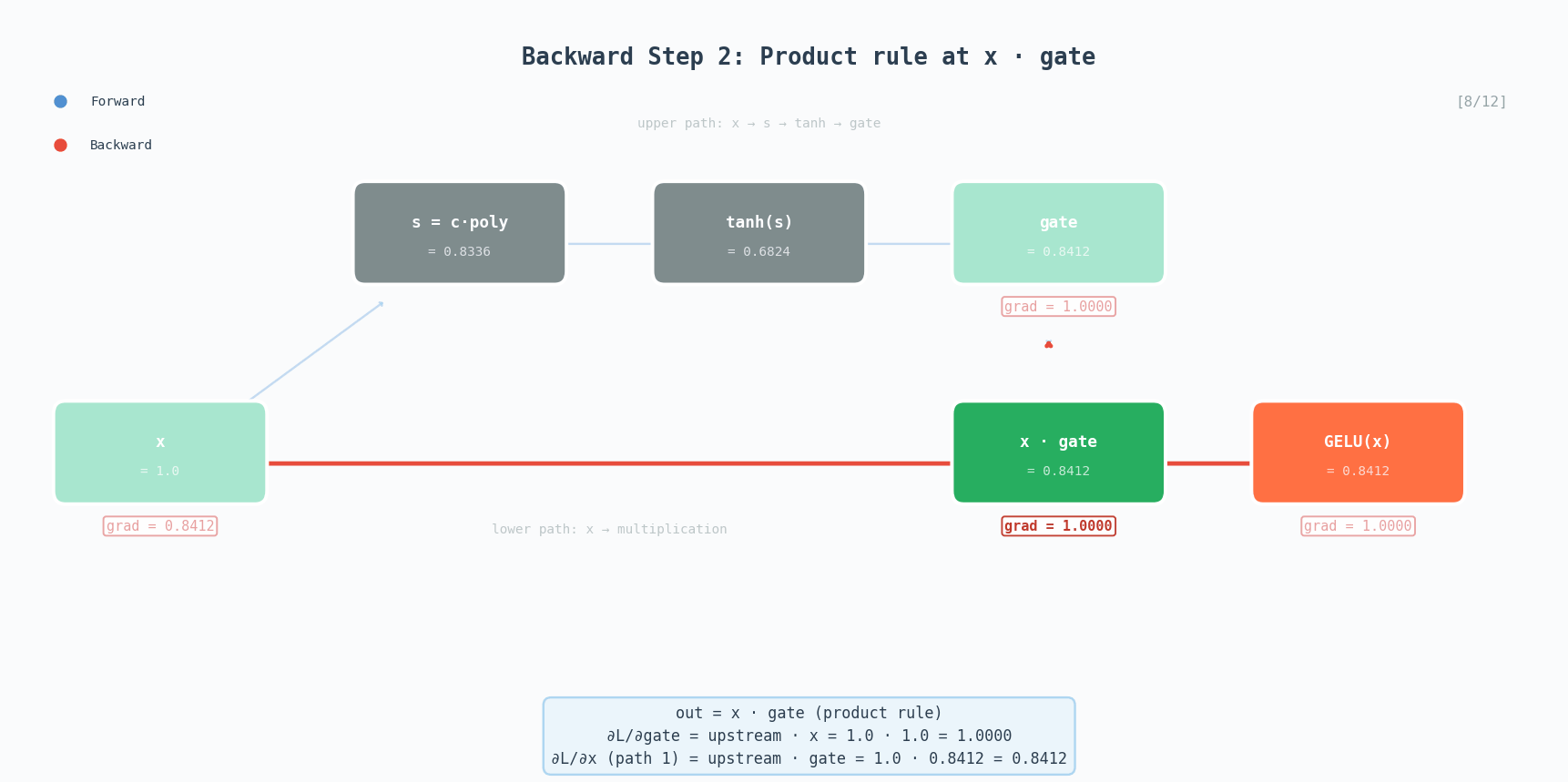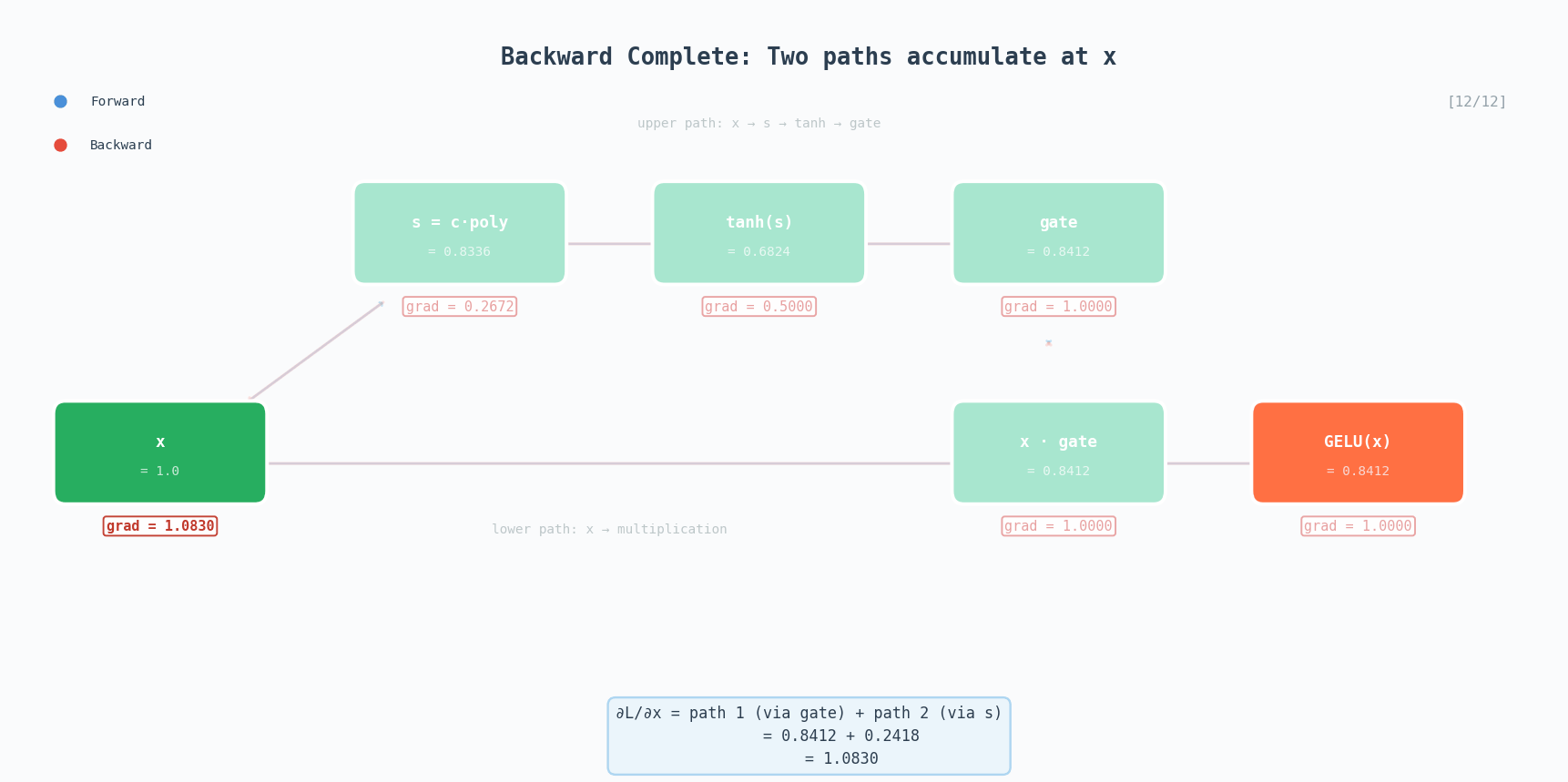
\[\text{GELU}(x) \approx 0.5x\left(1 + \tanh\left(\sqrt{\frac{2}{\pi}}\left(x + 0.044715x^3\right)\right)\right)\]GELU is the activation used in GPT-2, BERT, and most modern transformers. Unlike ReLU which has a hard cutoff at zero, GELU smoothly gates values, small negative inputs get attenuated rather than completely zeroed out.
"""
deeplygrad.nn: Neural network building blocks
"""
class GELU(Module):
def __init__(self):
super().__init__()
def forward(self, t: Tensor) -> Tensor:
c = xp.sqrt(2.0 / xp.pi)
x = t.data
inner = c * (x + 0.044715 * x ** 3)
tanh_inner = xp.tanh(inner)
out_data = 0.5 * x * (1 + tanh_inner)
out = Tensor(out_data, requires_grad=t.requires_grad)
if out.requires_grad:
out._parents = [t]
def _backward(grad_output: np.ndarray) -> None:
sech2 = 1.0 - tan_inner**2
d_inner = c * (1.0 + 3.0 * 0.044715 * x ** 2)
g = grad_output * (0.5 * (1.0 + tanh_inner) + 0.5 * x * sech2 * d_inner)
t.grad = t.grad + g if t.grad is not None else g
out._grad_fn = _backward
return out
We’ll use GELU in Part 3 when we build the transformer. For now, ReLU is what we need for our MLP.
Why do you need a custom backward?
Unlike ReLU, we use a custom _backward here. Why? Because we don’t have a tanh op on our Tensor, and even if we did, the composed backward would create many intermediate tensors. The fused version computes the derivative in one shot. PyTorch does the same, nn.GELU has a custom CUDA kernel. This tells us an important design principle for designing our own modules:
-
Compose when you can. If your
forward()only uses existing differentiable Tensor ops, autograd handles backward for you. This is the common case for most Modules we write. -
Fuse when you must. Write a custom _backward when you need numerical stability (like CrossEntropyLoss below) or when the composed version would be too slow or require ops you haven’t implemented yet (like GELU needing tanh).
This is the same tradeoff PyTorch makes. Most nn.Module subclasses don’t define backward at all. The few that do (like CrossEntropyLoss, GELU, LayerNorm) fuse for stability or performance.
Cross-Entropy Loss
For classification, we need softmax to convert raw scores (logits) into probabilities, and cross-entropy to measure how far those probabilities are from the true labels.
Why fuse them?
If we computed softmax and cross-entropy separately:
probs = softmax(logits) # can have values like 1e-45
loss = -log(probs[target]) # log(1e-45) = -103, numerically a disaster, unstable learning
The fused version avoids this using the log-sum-exp trick:
\[-\text{log}(\text{softmax}(x)_k) = -x_k + \text{log}\sum_j\text{exp}(x_j)\]And for numerical stability,
\[\text{log}\sum_j\text{exp}(x_j) = \text{max}(x) + \text{log}\sum_j\text{exp}(x_j - \text{max}(x))\]Subtracting the max means we never exponentiate large numbers. This is exactly what PyTorch’s nn.CrossEntropyLoss does internally. And this is why we write a custom _backward here rather than composing from primitive ops.
"""
deeplygrad.nn: Neural network building blocks
"""
class CrossEntropyLoss(Module):
def __init__(self):
super().__init__()
def forward(self, logits: Tensor, targets: Tensor) -> Tensor:
batch_size = logits.shape[0]
t = targets.data.astype(int)
x = logits.data
x_max = np.max(axis=1, keepdims=True)
x_shifted = x - x_max
exp_shifted = xp.exp(x_shifted)
sum_exp = exp_shifted.sum(axis=1, keepdims=True)
log_sum_exp = x_max + xp.log(sum_exp)
log_probs = x - log_sum_exp
correct_log_probs = log_probs[xp.arange(batch_size), t]
loss_val = -correct_log_probs.mean()
loss = Tensor(loss_val, requires_grad=logits.requires_grad)
if out.requires_grad:
out._parents = [logits]
probs = exp_shifted / sum_exp
def _backward(grad_output):
# Compute softmax probabilities
probs = xp.exp(log_probs)
# Create one-hot encoding of targets
one_hot = xp.zeros_like(probs)
one_hot[xp.arange(batch_size), targets.data.astype(int)] = 1
# Gradient w.r.t logits
grad_logits = ((probs - one_hot) / batch_size) * grad_output
logits.grad = logits.grad + grad_logits if logits.grad is not None else grad_logits
loss._grad_fn = _backward
return loss
The gradient formula softmax - one_hot is one of the most elegant results in deep learning. If the model predicts the correct class with 90% confidence, the gradient for that class is $0.9−1.0=−0.1$ (push the logit up a bit). For a wrong class predicted at 10%, the gradient is $0.1−0=0.1$ (push it down). The magnitude is proportional to how wrong the prediction is.
Adam Optimizer
In Part 1 we used manual SGD optimizer. Now let’s build Adam (Kingma & Ba, 2014), which is what used most widely. Adam maintains running averages of both the gradient (first moment $m$) and the squared gradient (second moment $v$), with bias correction:
\[m_t = \beta_1 m_{t-1} + (1-\beta_1) g_t, \quad v_t = \beta_2 v_{t-1} + (1-\beta_2) g_t^2\] \[\hat{m}_t = \frac{m_t}{1-\beta_1^t}, \quad \hat{v}_t = \frac{v_t}{1-\beta_2^t}\] \[\theta_t = \theta_{t-1} - \alpha \frac{\hat{m}_t}{\sqrt{\hat{v}_t} + \epsilon}\]The intuition: $m$ gives momentum (smooths out noisy gradients), $v$ gives per-parameter learning rate adaptation (parameters with consistently large gradients get smaller steps).
"""
deeplygrad.optim: Optimization algorithms
"""
class Adam:
def __init__(self, parameters, lr=0.001, beta1=0.9, beta2=0.999, eps=1e-8):
self.parameters = parameters
self.lr = lr
self.beta1, self.beta2, self.eps = beta1, beta2, eps
self.t = 0
self.m = [xp.zeros_like(p.data) for p in parameters]
self.v = [xp.zeros_like(p.data) for p in parameters]
def step(self):
self.t += 1
for i, p in enumerate(self.parameters):
if p.grad is None:
continue
self.m[i] = self.beta1 * self.m[i] + (1 - self.beta1) * p.grad
self.v[i] = self.beta2 * self.v[i] + (1 - self.beta2) * (p.grad ** 2)
m_hat = self.m[i] / (1 - self.beta1 ** self.t)
v_hat = self.v[i] / (1 - self.beta2 ** self.t)
p.data -= self.lr * m_hat / (xp.sqrt(v_hat) + self.eps)
Putting It All Together: MLP on Digit Recognition
Finally, We’ll train a neural network to classify MNIST digits using the deeplygrad library.
"""
deeplygrad.neural_network.mnist.py : Train a MLP on MNIST using the deeplygrad autograd engine
"""
import sys, os
from torchvision.datasets import MNIST
from deeplygrad.nn import Module, Linear, ReLU, CrossEntropyLoss
from deeplygrad.optim import Adam
from deeplygrad.backend import xp, BACKEND_NAME
from deeplygrad.tensor import Tensor
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
import numpy as np
print(f"Using backend: {BACKEND_NAME}")
def get_data():
train_dataset = MNIST(root='./data', train=True, download=True)
test_dataset = MNIST(root='./data', train=False, download=True)
return train_dataset, test_dataset
def preprocess_data(dataset):
images = dataset.data
labels = dataset.targets
images = images.reshape(-1, 28 * 28) / 255.0
labels = labels.long()
return images, labels
class MLP(Module):
"""
3-layer MLP: 784 -> 128 -> 64 -> 10
Activation: ReLU
Loss: Cross-Entropy Loss
Optimizer: Adam
"""
def __init__(self):
super().__init__()
self.fc1 = Linear(784, 128)
self.relu1 = ReLU()
self.fc2 = Linear(128, 64)
self.relu2 = ReLU()
self.fc3 = Linear(64, 10)
def forward(self, x):
x = self.fc1(x)
x = self.relu1(x)
x = self.fc2(x)
x = self.relu2(x)
x = self.fc3(x)
return x
def main():
xp.random.seed(42)
train_dataset, test_dataset = get_data()
train_images, train_labels = preprocess_data(train_dataset)
test_images, test_labels = preprocess_data(test_dataset)
model = MLP()
optimizer = Adam(model.parameters(), lr=0.001)
criterion = CrossEntropyLoss()
# Split: 8000 train, 2000 val from training set; test set separate
val_images = train_images[8000:10000]
val_labels = train_labels[8000:10000]
train_images = train_images[:8000]
train_labels = train_labels[:8000]
test_images = test_images[:1000]
test_labels = test_labels[:1000]
batch_size = 32
epochs = 10
epoch_train_losses = []
epoch_val_losses = []
batch_losses = []
test_images_t = Tensor(test_images.numpy(), requires_grad=False)
test_labels_t = Tensor(test_labels.numpy(), requires_grad=False)
val_images_t = Tensor(val_images.numpy(), requires_grad=False)
val_labels_t = Tensor(val_labels.numpy(), requires_grad=False)
for epoch in range(epochs):
epoch_batch_losses = []
for i in range(0, len(train_images), batch_size):
images = train_images[i:i+batch_size].numpy()
labels = train_labels[i:i+batch_size].numpy()
images = Tensor(images, requires_grad=False)
labels = Tensor(labels, requires_grad=False)
optimizer.zero_grad()
outputs = model(images)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
l = loss.item()
epoch_batch_losses.append(l)
batch_losses.append(l)
avg_train_loss = np.mean(epoch_batch_losses)
epoch_train_losses.append(avg_train_loss)
val_outputs = model(val_images_t)
val_loss = criterion(val_outputs, val_labels_t).item()
epoch_val_losses.append(val_loss)
val_preds = xp.argmax(val_outputs.data, axis=1)
val_accuracy = float(xp.mean((val_preds == val_labels_t.data).astype(float)))
print(f"Epoch {epoch+1}/{epochs}, Train Loss: {avg_train_loss:.4f}, "
f"Val Loss: {val_loss:.4f}, Val Acc: {val_accuracy:.4f}")
print("Training complete")
test_outputs = model(test_images_t)
test_preds = xp.argmax(test_outputs.data, axis=1)
test_accuracy = float(xp.mean((test_preds == test_labels_t.data).astype(float)))
print(f"Final test accuracy: {test_accuracy:.4f}")
main()
Using backend: numpy
Epoch 1/10, Train Loss: 0.5647, Val Loss: 0.3684, Val Acc: 0.8905
Epoch 2/10, Train Loss: 0.2366, Val Loss: 0.2881, Val Acc: 0.9195
Epoch 3/10, Train Loss: 0.1548, Val Loss: 0.2600, Val Acc: 0.9255
Epoch 4/10, Train Loss: 0.1068, Val Loss: 0.2427, Val Acc: 0.9295
Epoch 5/10, Train Loss: 0.0765, Val Loss: 0.2300, Val Acc: 0.9320
Epoch 6/10, Train Loss: 0.0550, Val Loss: 0.2367, Val Acc: 0.9330
Epoch 7/10, Train Loss: 0.0407, Val Loss: 0.2363, Val Acc: 0.9360
Epoch 8/10, Train Loss: 0.0318, Val Loss: 0.2362, Val Acc: 0.9400
Epoch 9/10, Train Loss: 0.0254, Val Loss: 0.2752, Val Acc: 0.9335
Epoch 10/10, Train Loss: 0.0212, Val Loss: 0.2694, Val Acc: 0.9340
Training complete
Final test accuracy: 0.9360
This is structurally identical to a PyTorch training loop. model(x) calls forward(), criterion(logits, targets) computes the loss, optimizer.zero_grad() resets all gradients, loss.backward() propagates them, and optimizer.step() updates the weights. Every operation, the matmul, the ReLU, the cross-entropy, the Adam update, is code we wrote ourselves.
What We Built in Part 2
All built on the Tensor from Part 1, everything inheriting from Module:
-
Module: The base class with__init__,parameters(),zero_grad(),train()/eval(), andregister_buffer(). -
Linear: Fully connected layer with Kaiming initialization. -
ReLU: Composed from existing Tensor ops (no custom backward needed). -
GELU: Custom backward for efficiency (we’ll use this for the transformer). -
CrossEntropyLoss: Custom backward for numerical stability (log-sum-exp trick). -
Adam: Adaptive optimizer with momentum and per-parameter learning rates.
Key Takeaways
- Everything is a Module. Layers, activations, losses. They snap together like LEGO blocks, composing into arbitrarily complex architectures.
- Compose when you can, fuse when you must. ReLU uses existing Tensor ops and gets backward for free. CrossEntropyLoss and GELU write custom backward for stability and performance.
-
backward()lives onTensor, notModule. Modules organize the forward pass. Tensors own the computation graph.loss.backward()walks the Tensor graph directly. - Nonlinear activations are what make deep learning deep. Without them, any stack of linear layers collapses to a single linear transformation.
- The gradient of cross-entropy w.r.t. logits is
softmax - one_hot, simple and proportional to how wrong the prediction is. - The training loop pattern (forward, loss, zero_grad, backward, step) is universal. Our from-scratch version is structurally identical to PyTorch.
What’s Next
In Part 3, we build a transformer. We’ll implement multi-head attention, layer normalization, positional encoding, and the full transformer block, then train a character-level language model. Everything built on the autograd engine from Parts 1 and 2. The full code is available in the simplygrad repository.
Related Posts
Here are some more articles you might like to read next: